Functional data for any clinically relevant gene variant.
geneSlice® is our proprietary gene editing platform. It generates the precise, uniform variant libraries — and single-variant edits — that high-throughput functional assays require, so we can classify every variant in a target gene accurately and within drug development timelines.

Our proprietary gene editing platform — precise, uniform variant libraries and single-variant edits, built for high-throughput functional assays.
Serine integrase-based gene editing
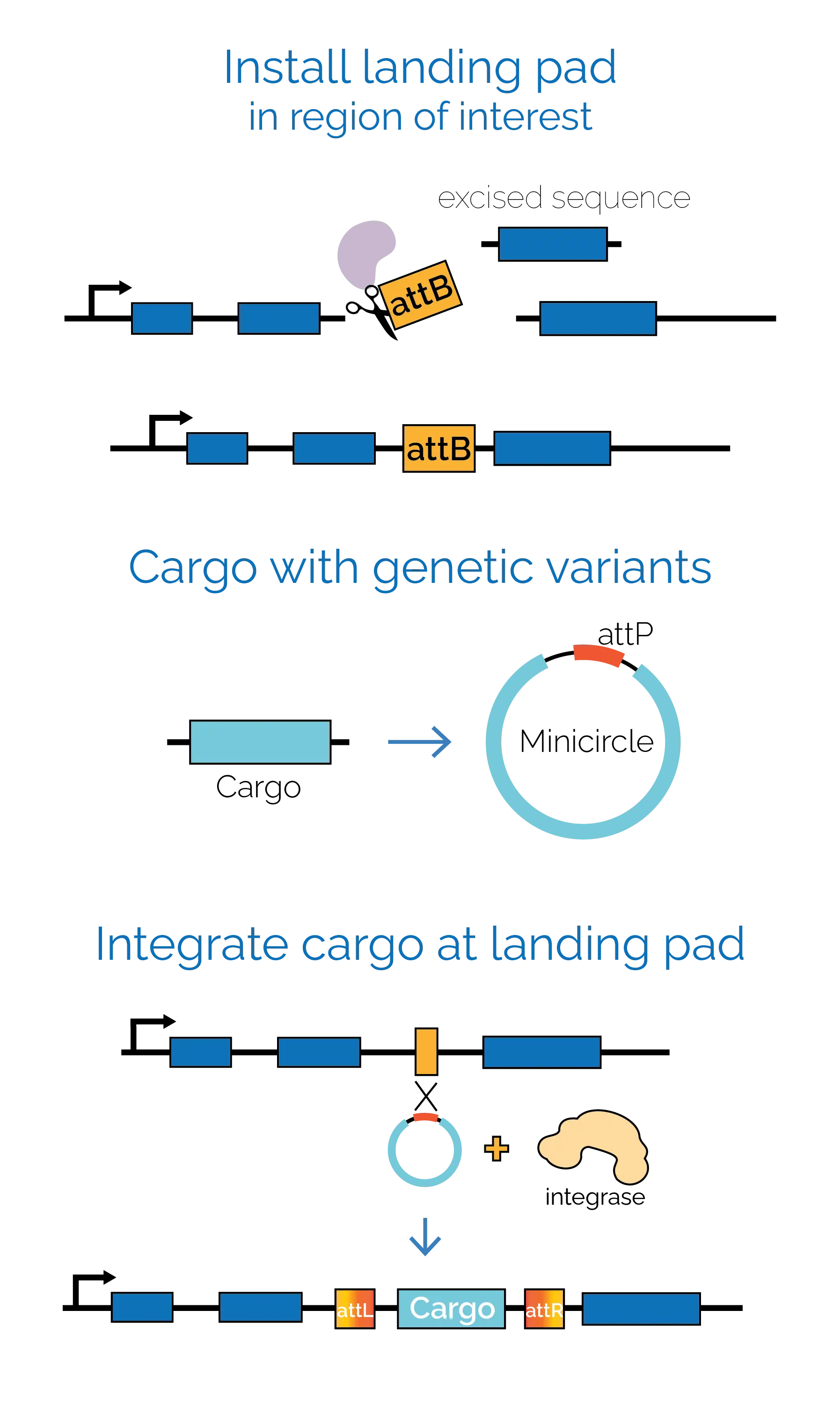
geneSlice is built on serine integrase-based gene editing — a mature class of bacteriophage enzymes that catalyse precise, unidirectional DNA recombination at defined sequences. The platform decouples where a variant is introduced from what the variant is, in a two-step approach:
- Install a landing pad in a defined location in the cell genome. This step is performed once per cell line.
- Deliver variant cassettes to the landing pad using serine integrase recombination. Cassettes can be introduced as single variants for targeted edits, or as full libraries in which each cell carries a different gene variant at the same genomic position.
We integrate variants without perturbing the target gene's expression. Our in-house computational design pipeline combines algorithms that position the landing pad and select integration scars to leave the endogenous locus undisturbed — a property that makes geneSlice especially well suited to assays where preserving native expression matters.
geneSlice can also tag proteins of interest with various tags to support your workflow — including fluorescent proteins (GFP, mNeonGreen, mScarlet), epitope tags (FLAG, HA, V5, Myc), and luminescent reporters (HiBiT, NanoLuc).
What this enables
Saturation mutagenesis
Across user-defined regions of interest.
Single-variant precision edits
For targeted programmes.
Complex variants
Large insertions and deletions, splice-site and regulatory variants, structural changes.
Difficult genomic contexts
Repetitive sequences, highly homologous regions, and loci with high off-target liability.
Gene tagging
At the endogenous locus — fluorescent, epitope, and luminescent tags.
CRISPR therapeutics characterisation
Mapping bystander mutations, off-target effects, and unintended edits introduced by therapeutic editing tools.
How geneSlice compares
CRISPR HDR, prime editing, and base editing each have sweet spots — short edits, clean small variants, single-nucleotide transitions. None handles the full variant space at scale, especially in repetitive or highly homologous regions. geneSlice fills that gap. By separating variant delivery from variant content, we introduce any variant — short or long, simple or complex — uniformly at a defined locus.
| Feature | geneSlice | Prime Editing | Cas9 HDR | Base Editing |
|---|---|---|---|---|
| ROI precision | High | High | Variable | Limited window |
| Repetitive regions | ✓ | ✗ | ✗ | ✗ |
| Homologous regions | ✓ | ~ | ✗ | ~ |
| Off-target risk | Low | Low | High | Medium |
| Saturation mutagenesis | Full ROI | Limited | ✗ | Limited |
| Complex edits | ✓ | ✓ | ~ | �✗ |
From editing to functional data
geneSlice projects run on one of two pathways.
Variant library workflow
The productised pipeline. Best fit for saturation mutagenesis, large-scale variant classification, and trial-enabling variant databases.
Consultation
Define gene, region of interest, variant library, phenotypic readout.
2–4 weeksCell line development
Install landing pad in a disease-relevant cell line.
~6 monthsLibrary & screening
Generate cassette library, deliver via geneSlice, run functional assay.
~6 monthsData delivery
Functional scores per variant, resistance maps, raw data on request.
deliverableBespoke project workflow
Custom assays where the standard library pipeline isn't the right fit — single-variant studies, tagging-based engagement assays, custom phenotypic endpoints, characterisation work for CRISPR therapeutics, and others.
Consultation
Define scientific question and assay architecture.
2–4 weeksBespoke assay design
Custom cell-line build, custom edit strategy, custom readouts.
scopedExecution
Functional assays, arrayed screens, drug sensitivity assays, with bioinformatic support throughout.
scopedData delivery
As agreed.
deliverableTypical Path 1 projects run 6–12 months end-to-end depending on scope. Path 2 timelines scope per project.
What clients receive
Deliverables vary by project and are agreed up front:
- Functional scores per variant — the core data product.
- Resistance and sensitivity maps for compound–target programmes.
- Annotated variant databases suitable for trial recruitment workflows.
- Tagged cell lines and engagement-assay data for bespoke projects involving tagging.
- Raw data, analysis pipelines, and reports as agreed.
geneSlice® is patented, with exclusive license held by VUS Genetics.