Functional answers for rare disease programmes — from diagnosis to therapeutic response.
Disease-relevant cell models and functional assays across rare disease, neurodegenerative, cardiovascular, and other genetically defined disorders.
For rare genetic diseases, drug developers face two functional questions at once: which variants drive disease, and which patients will respond to a candidate therapy. We use geneSlice to build disease-relevant cell models carrying patient-specific variants, then measure their phenotypic consequence and therapeutic response — supporting variant classification and clinical trial design across rare genetic disease, neurodegenerative, cardiovascular, and other genetically defined disorders.
Every patient counts.
In rare genetic disease, trial populations are small, many variants are private to single families, and the cost of mis-stratifying patients — enrolling non-responders or excluding likely responders — is disproportionately high. Variant-level functional data resolves this: it confirms which variants are truly pathogenic, separates loss-of-function from partial-function, and predicts how a candidate therapy will perform across the variant spectrum, before clinical exposure.
From diagnosis to therapeutic response
Variant classification for diagnosis & trial design
Functional confirmation of pathogenicity for variants of uncertain significance. Supports patient stratification for clinical trials, and provides foundational data for variant reclassification in genetic diagnostic workflows.
Therapeutic response prediction
For drug developers, we test how candidate therapies perform across the variant spectrum — predicting which patients are most likely to benefit, and identifying variants where the therapy may not reverse disease phenotype.
Custom cell models & assays
For projects outside the standard variant-classification shape — bespoke phenotypic readouts, complex variant combinations, tagging-based readouts — see Custom Services.
Niemann-Pick type C (NPC)
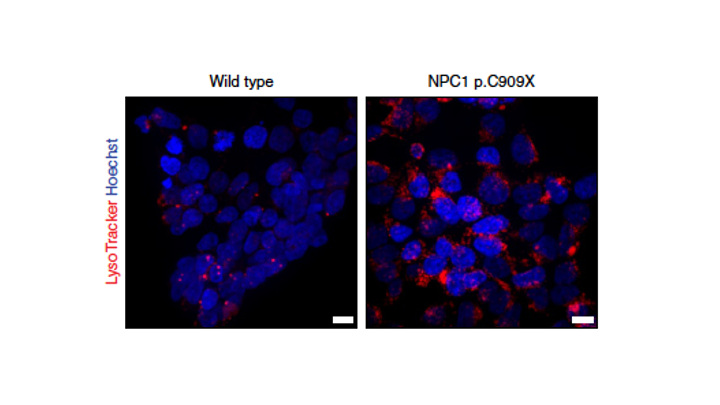
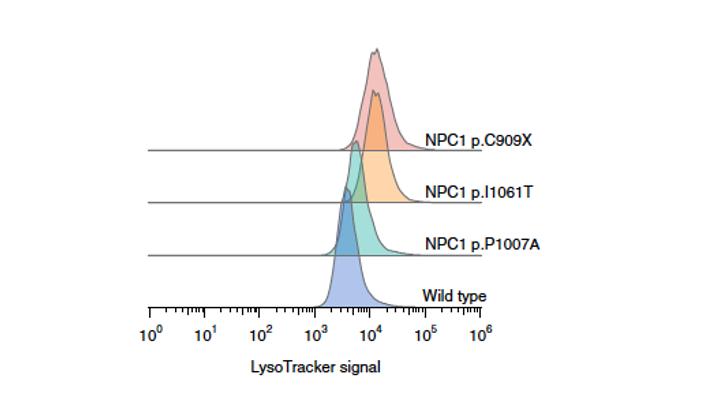
NPC is a rare neurological disorder caused by loss-of-function variants in NPC1, which encodes an intracellular cholesterol transporter. Pathogenic variants drive cholesterol accumulation in lysosomes.
Team member Zhenya Ivakine established a functional readout for the disease — quantifying lysosomal cholesterol via LysoTracker — and scored thousands of NPC1 variants for functional impact. The result is a variant-level functional map distinguishing loss-of-function from functional variants with high specificity.
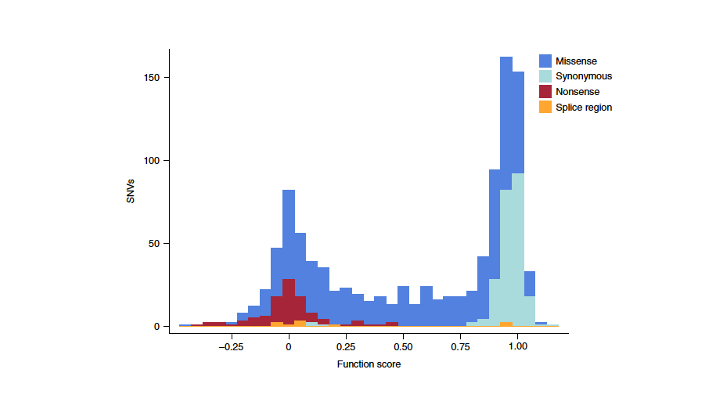
This kind of analysis turns uncertain diagnoses into actionable classifications, and gives drug developers a quantitative basis for predicting which patients will respond to NPC1-targeted therapies.
Standard geneSlice pipeline
Consultation → cell line development (~6 mo) → variant library and screening (~6 mo) → data delivery. Typically 6–12 months. See Technology.